Sanger sequencing, also known as chain-termination sequencing, refers to a method of DNA sequencing developed by Frederick Sanger in 1977. This method is based on amplification of the DNA fragment to be sequenced by DNA polymerase and incorporation of modified nucleotides - specifically, dideoxynucleotides (ddNTPs).
The classical chain-termination method requires a single-stranded DNA template, a DNA primer, a DNA polymerase, normal deoxynucleotidetriphosphates (dNTPs), and modified nucleotides (dideoxyNTPs) that terminate DNA strand elongation . These chain-terminating nucleotides lack a 3'-OH group required for the formation of a phosphodiester bond between two nucleotides, causing DNA polymerase to cease extension of DNA when a ddNTP is incorporated. The ddNTPs may be radioactively or fluorescently labelled for detection in automated sequencing machines.The DNA sample is divided into four separate sequencing reactions, containing all four of the standard deoxynucleotides (dATP, dGTP, dCTP and dTTP) and the DNA polymerase. To each reaction is added only one of the four dideoxynucleotides (ddATP, ddGTP, ddCTP, or ddTTP). Following rounds of template DNA extension from the bound primer, the resulting DNA fragments are heat denatured and separated by size using gel electrophoresis. This is frequently performed using a denaturing polyacrylamide-urea gel with each of the four reactions run in one of four individual lanes (lanes A, T, G, C). The DNA bands may then be visualized by autoradiography or UV light and the DNA sequence can be directly read off the X-ray film or gel image.
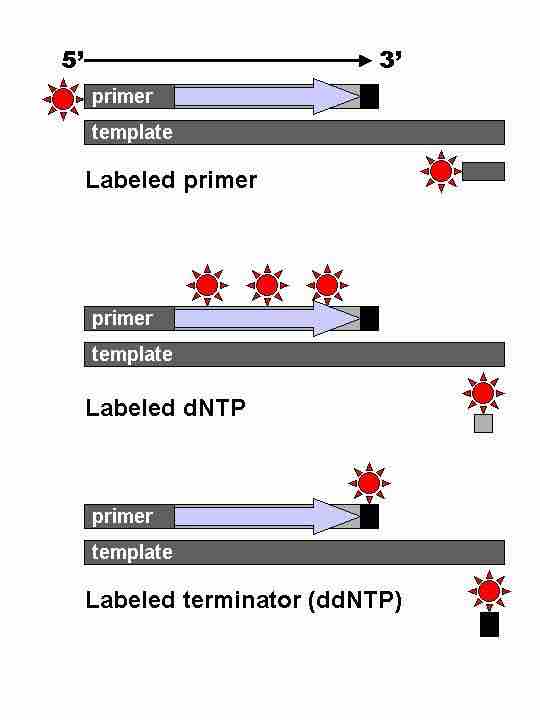
Sanger sequencing
Different types of Sanger sequencing, all of which depend on the sequence being stopped by a terminating dideoxynucleotide (black bars).
Technical variations of chain-termination sequencing include tagging with nucleotides containing radioactive phosphorus for radiolabelling, or using a primer labeled at the 5' end with a fluorescent dye. Dye-primer sequencing facilitates reading in an optical system for faster and more economical analysis and automation. The later development by Leroy Hood and coworkers of fluorescently labeled ddNTPs and primers set the stage for automated, high-throughput DNA sequencing. Chain-termination methods have greatly simplified DNA sequencing. More recently, dye-terminator sequencing has been developed. Dye-terminator sequencing utilizes labelling of the chain terminator ddNTPs, which permits sequencing in a single reaction, rather than four reactions as in the labelled-primer method. In dye-terminator sequencing, each of the four dideoxynucleotide chain terminators is labelled with fluorescent dyes, each of which emit light at different wavelengths .
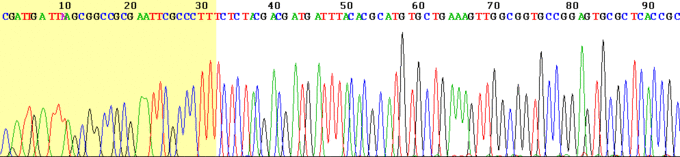
Chromatograph
This is an example of the output of a Sanger sequencing read using fluorescently labelled dye-terminators. The four DNA bases are represented by different colours which are interpreted by the software to give the DNA sequence above.
Automated DNA-sequencing instruments (DNA sequencers) can sequence up to 384 DNA samples in a single batch (run) in up to 24 runs a day. DNA sequencers carry out capillary electrophoresis for size separation, detection and recording of dye fluorescence, and data output as fluorescent peak trace chromatograms. Automation has lead to the sequencing of entire genomes.